Plotting¶
Introduction¶
Labeled data enables expressive computations. These same labels can also be used to easily create informative plots.
xarray’s plotting capabilities are centered around
xarray.DataArray objects.
To plot xarray.Dataset objects
simply access the relevant DataArrays, ie dset['var1'].
Here we focus mostly on arrays 2d or larger. If your data fits
nicely into a pandas DataFrame then you’re better off using one of the more
developed tools there.
xarray plotting functionality is a thin wrapper around the popular matplotlib library. Matplotlib syntax and function names were copied as much as possible, which makes for an easy transition between the two. Matplotlib must be installed before xarray can plot.
For more extensive plotting applications consider the following projects:
- Seaborn: “provides a high-level interface for drawing attractive statistical graphics.” Integrates well with pandas.
- Holoviews: “Composable, declarative data structures for building even complex visualizations easily.” Works for 2d datasets.
- Cartopy: Provides cartographic tools.
Imports¶
The following imports are necessary for all of the examples.
In [1]: import numpy as np
In [2]: import pandas as pd
In [3]: import matplotlib.pyplot as plt
In [4]: import xarray as xr
For these examples we’ll use the North American air temperature dataset.
In [5]: airtemps = xr.tutorial.load_dataset('air_temperature')
In [6]: airtemps
Out[6]:
<xarray.Dataset>
Dimensions: (lat: 25, lon: 53, time: 2920)
Coordinates:
* lat (lat) float32 75.0 72.5 70.0 67.5 65.0 62.5 60.0 57.5 55.0 52.5 ...
* time (time) datetime64[ns] 2013-01-01 2013-01-01T06:00:00 ...
* lon (lon) float32 200.0 202.5 205.0 207.5 210.0 212.5 215.0 217.5 ...
Data variables:
air (time, lat, lon) float64 241.2 242.5 243.5 244.0 244.1 243.9 ...
Attributes:
platform: Model
Conventions: COARDS
references: http://www.esrl.noaa.gov/psd/data/gridded/data.ncep.reanalysis.html
description: Data is from NMC initialized reanalysis
(4x/day). These are the 0.9950 sigma level values.
title: 4x daily NMC reanalysis (1948)
# Convert to celsius
In [7]: air = airtemps.air - 273.15
One Dimension¶
Simple Example¶
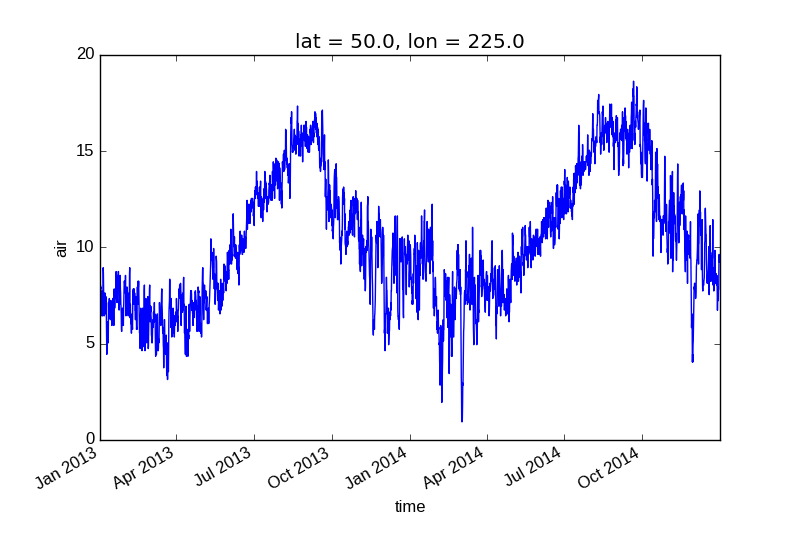
xarray uses the coordinate name to label the x axis.
In [8]: air1d = air.isel(lat=10, lon=10)
In [9]: air1d.plot()
Out[9]: [<matplotlib.lines.Line2D at 0x7f08043e5b50>]
Additional Arguments¶
Additional arguments are passed directly to the matplotlib function which
does the work.
For example, xarray.plot.line() calls
matplotlib.pyplot.plot passing in the index and the array values as x and y, respectively.
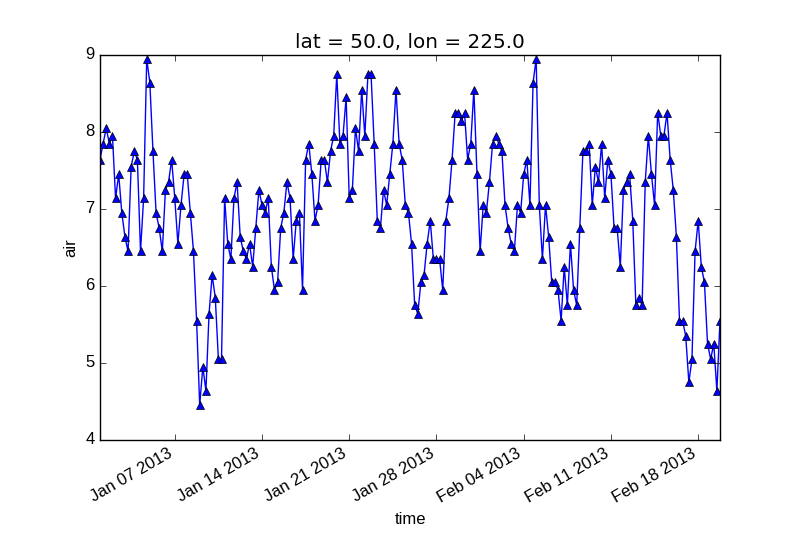
So to make a line plot with blue triangles a matplotlib format string
can be used:
In [10]: air1d[:200].plot.line('b-^')
Out[10]: [<matplotlib.lines.Line2D at 0x7f07d8561290>]
Note
Not all xarray plotting methods support passing positional arguments to the wrapped matplotlib functions, but they do all support keyword arguments.
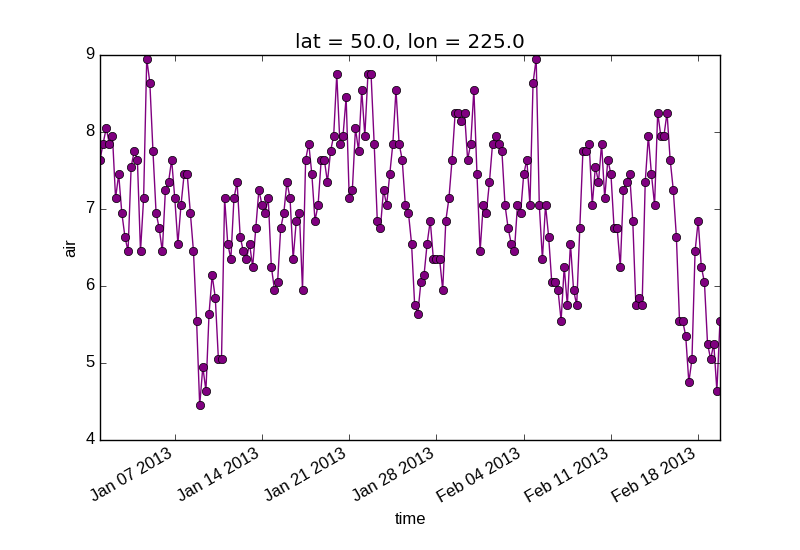
Keyword arguments work the same way, and are more explicit.
In [11]: air1d[:200].plot.line(color='purple', marker='o')
Out[11]: [<matplotlib.lines.Line2D at 0x7f07d854bfd0>]
Adding to Existing Axis¶
To add the plot to an existing axis pass in the axis as a keyword argument
ax. This works for all xarray plotting methods.
In this example axes is an array consisting of the left and right
axes created by plt.subplots.
In [12]: fig, axes = plt.subplots(ncols=2)
In [13]: axes
Out[13]:
array([<matplotlib.axes._subplots.AxesSubplot object at 0x7f07e6195e90>,
<matplotlib.axes._subplots.AxesSubplot object at 0x7f07e6195f90>], dtype=object)
In [14]: air1d.plot(ax=axes[0])
Out[14]: [<matplotlib.lines.Line2D at 0x7f07e629c8d0>]
In [15]: air1d.plot.hist(ax=axes[1])
Out[15]:
(array([ 9., 38., 255., 584., 542., 489., 368., 258., 327., 50.]),
array([ 0.95 , 2.719, 4.488, ..., 15.102, 16.871, 18.64 ]),
<a list of 10 Patch objects>)
In [16]: plt.tight_layout()
In [17]: plt.show()
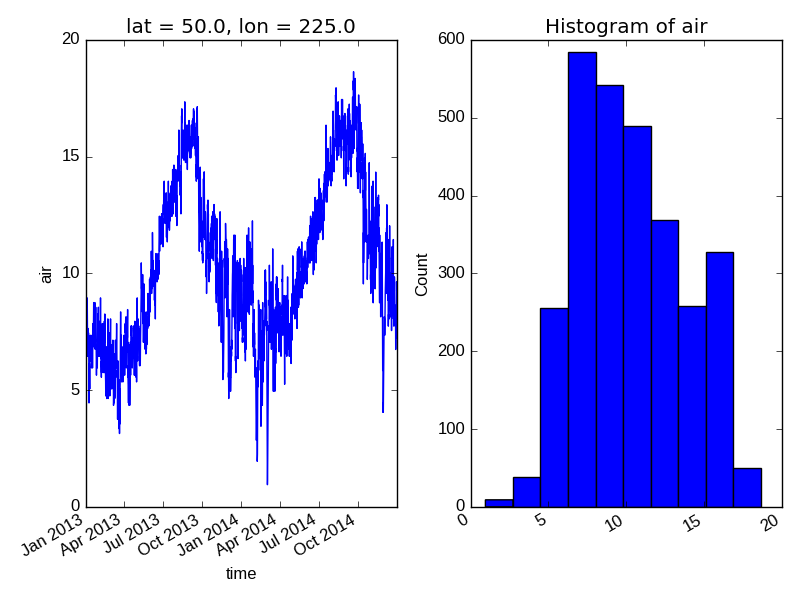
On the right is a histogram created by xarray.plot.hist().
Two Dimensions¶
Simple Example¶
The default method xarray.DataArray.plot() sees that the data is
2 dimensional and calls xarray.plot.pcolormesh().
In [18]: air2d = air.isel(time=500)
In [19]: air2d.plot()
Out[19]: <matplotlib.collections.QuadMesh at 0x7f07e6c942d0>
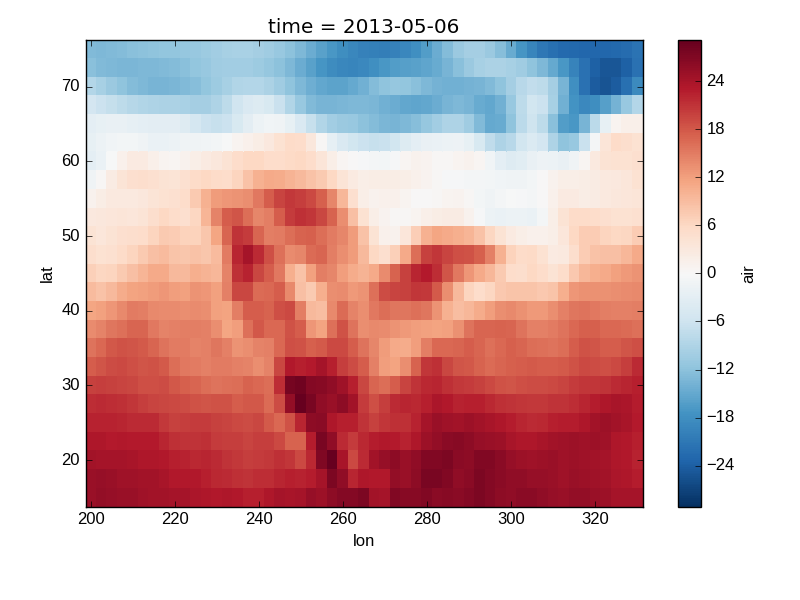
All 2d plots in xarray allow the use of the keyword arguments yincrease
and xincrease.
In [20]: air2d.plot(yincrease=False)
Out[20]: <matplotlib.collections.QuadMesh at 0x7f07e40fd1d0>

Note
We use xarray.plot.pcolormesh() as the default two-dimensional plot
method because it is more flexible than xarray.plot.imshow().
However, for large arrays, imshow can be much faster than pcolormesh.
If speed is important to you and you are plotting a regular mesh, consider
using imshow.
Missing Values¶
xarray plots data with Missing values.
In [21]: bad_air2d = air2d.copy()
In [22]: bad_air2d[dict(lat=slice(0, 10), lon=slice(0, 25))] = np.nan
In [23]: bad_air2d.plot()
Out[23]: <matplotlib.collections.QuadMesh at 0x7f07d81c7490>

Nonuniform Coordinates¶
It’s not necessary for the coordinates to be evenly spaced. Both
xarray.plot.pcolormesh() (default) and xarray.plot.contourf() can
produce plots with nonuniform coordinates.
In [24]: b = air2d.copy()
# Apply a nonlinear transformation to one of the coords
In [25]: b.coords['lat'] = np.log(b.coords['lat'])
In [26]: b.plot()
Out[26]: <matplotlib.collections.QuadMesh at 0x7f07d83aaed0>
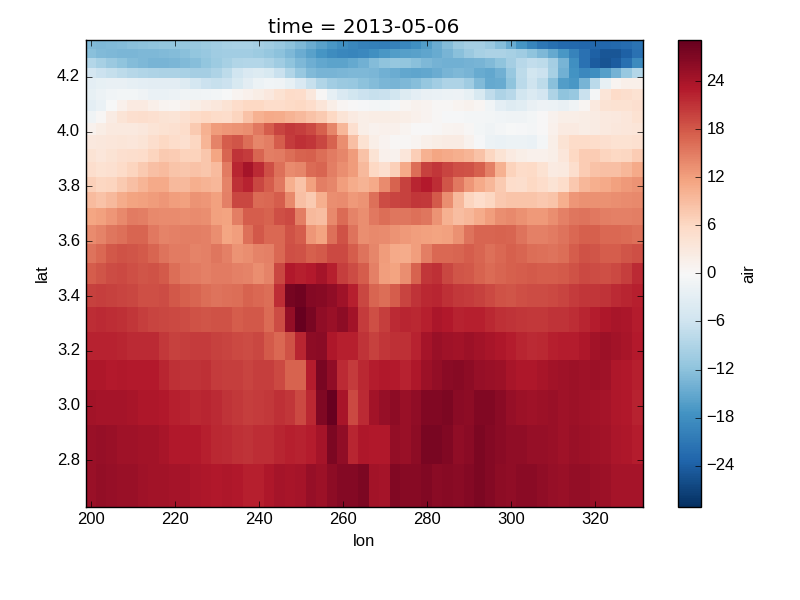
Calling Matplotlib¶
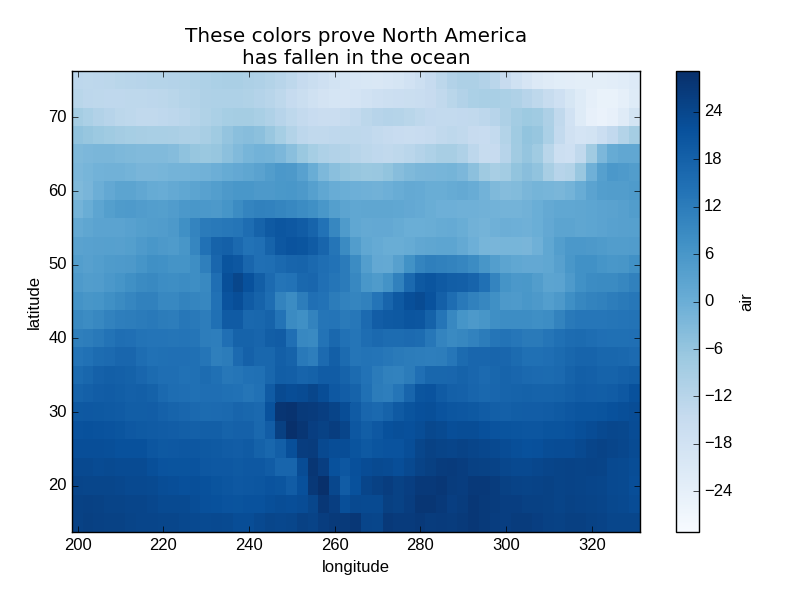
Since this is a thin wrapper around matplotlib, all the functionality of matplotlib is available.
In [27]: air2d.plot(cmap=plt.cm.Blues)
Out[27]: <matplotlib.collections.QuadMesh at 0x7f07d77d5150>
In [28]: plt.title('These colors prove North America\nhas fallen in the ocean')
Out[28]: <matplotlib.text.Text at 0x7f07d7810a50>
In [29]: plt.ylabel('latitude')
Out[29]: <matplotlib.text.Text at 0x7f07d78e1f90>
In [30]: plt.xlabel('longitude')
Out[30]: <matplotlib.text.Text at 0x7f07d78a4610>
In [31]: plt.tight_layout()
In [32]: plt.show()
Note
xarray methods update label information and generally play around with the
axes. So any kind of updates to the plot
should be done after the call to the xarray’s plot.
In the example below, plt.xlabel effectively does nothing, since
d_ylog.plot() updates the xlabel.
In [33]: plt.xlabel('Never gonna see this.')
Out[33]: <matplotlib.text.Text at 0x7f07d775c310>
In [34]: air2d.plot()
Out[34]: <matplotlib.collections.QuadMesh at 0x7f07e7848a10>
In [35]: plt.show()
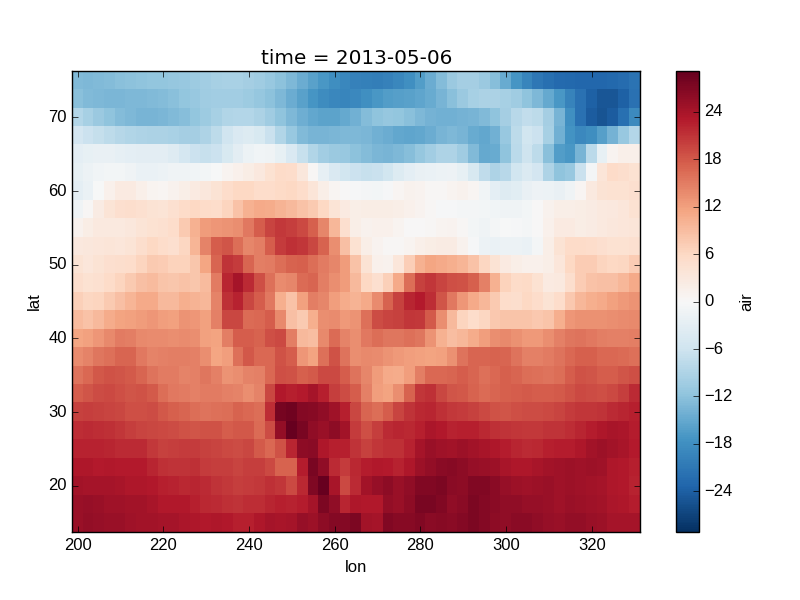
Colormaps¶
xarray borrows logic from Seaborn to infer what kind of color map to use. For example, consider the original data in Kelvins rather than Celsius:
In [36]: airtemps.air.isel(time=0).plot()
Out[36]: <matplotlib.collections.QuadMesh at 0x7f07d76c05d0>
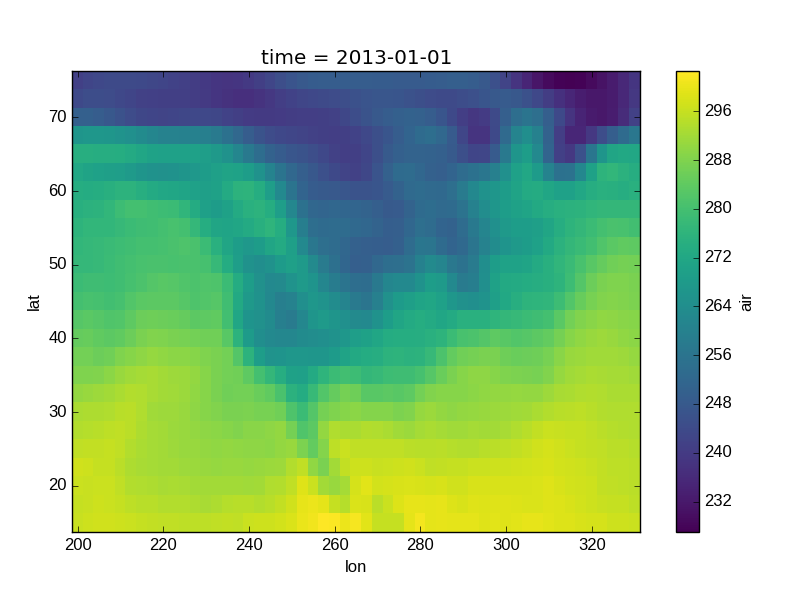
The Celsius data contain 0, so a diverging color map was used. The Kelvins do not have 0, so the default color map was used.
Robust¶
Outliers often have an extreme effect on the output of the plot. Here we add two bad data points. This affects the color scale, washing out the plot.
In [37]: air_outliers = airtemps.air.isel(time=0).copy()
In [38]: air_outliers[0, 0] = 100
In [39]: air_outliers[-1, -1] = 400
In [40]: air_outliers.plot()
Out[40]: <matplotlib.collections.QuadMesh at 0x7f07d77f0d50>
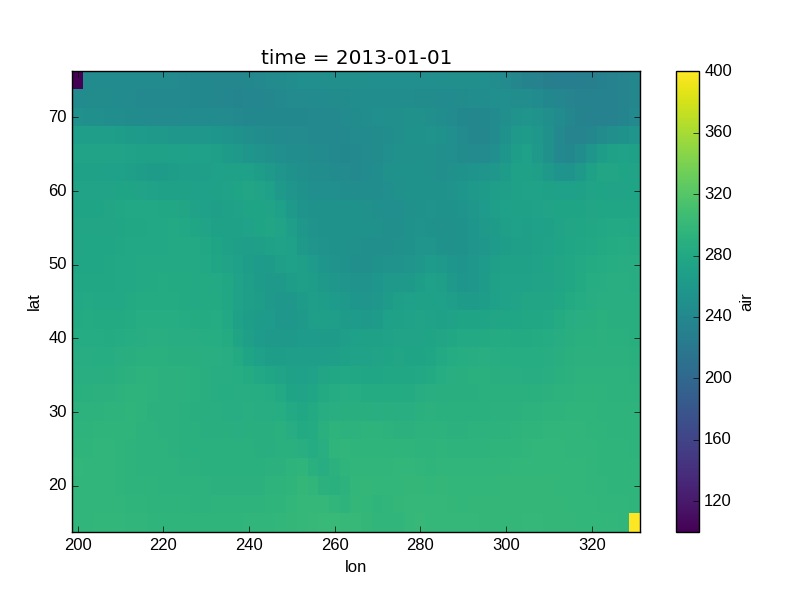
This plot shows that we have outliers. The easy way to visualize
the data without the outliers is to pass the parameter
robust=True.
This will use the 2nd and 98th
percentiles of the data to compute the color limits.
In [41]: air_outliers.plot(robust=True)
Out[41]: <matplotlib.collections.QuadMesh at 0x7f07e79d28d0>
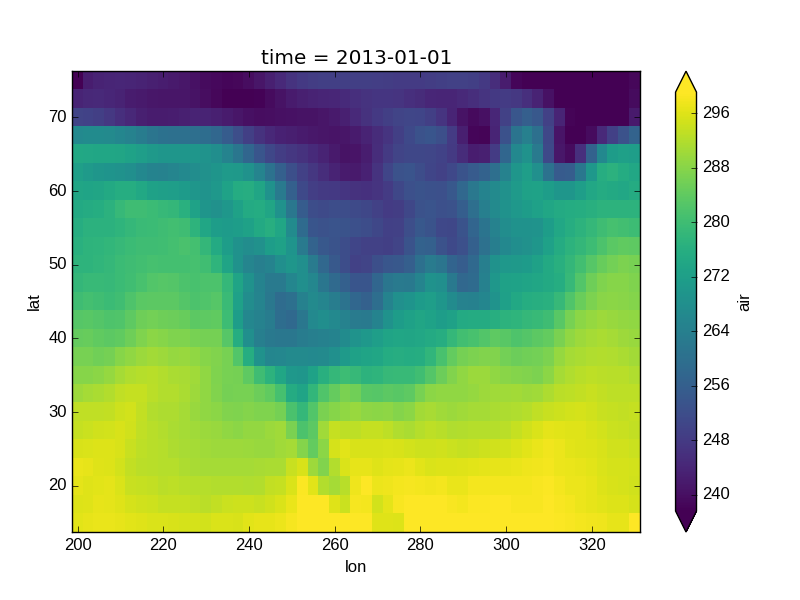
Observe that the ranges of the color bar have changed. The arrows on the color bar indicate that the colors include data points outside the bounds.
Discrete Colormaps¶
It is often useful, when visualizing 2d data, to use a discrete colormap,
rather than the default continuous colormaps that matplotlib uses. The
levels keyword argument can be used to generate plots with discrete
colormaps. For example, to make a plot with 8 discrete color intervals:
In [42]: air2d.plot(levels=8)
Out[42]: <matplotlib.collections.QuadMesh at 0x7f07d73a2850>
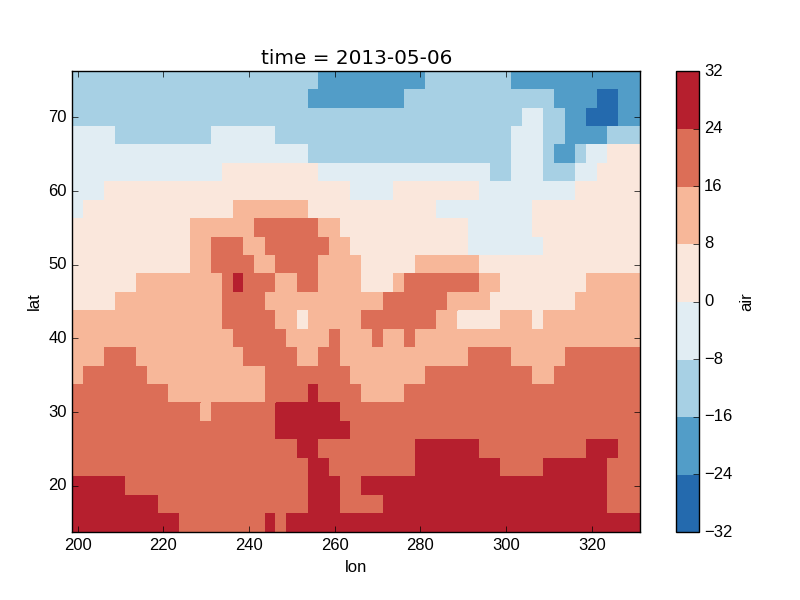
It is also possible to use a list of levels to specify the boundaries of the discrete colormap:
In [43]: air2d.plot(levels=[0, 12, 18, 30])
Out[43]: <matplotlib.collections.QuadMesh at 0x7f07d72a18d0>
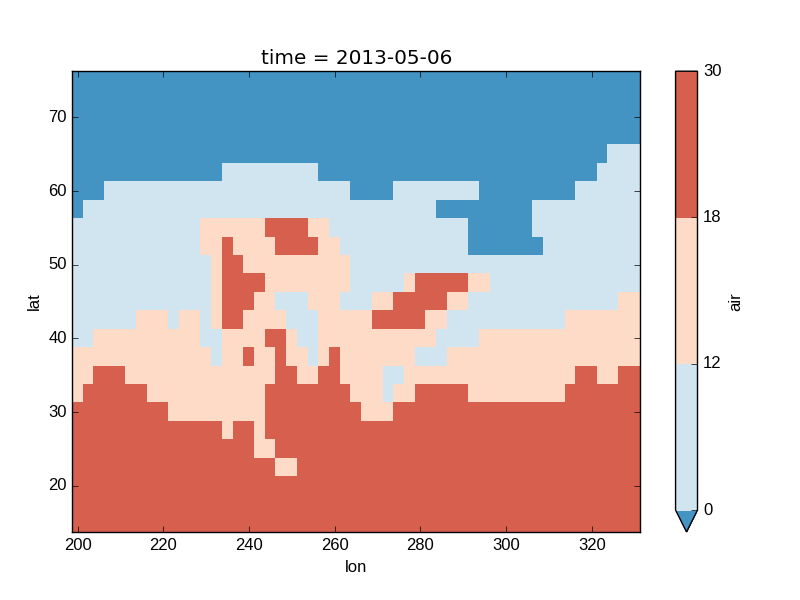
You can also specify a list of discrete colors through the colors argument:
In [44]: flatui = ["#9b59b6", "#3498db", "#95a5a6", "#e74c3c", "#34495e", "#2ecc71"]
In [45]: air2d.plot(levels=[0, 12, 18, 30], colors=flatui)
Out[45]: <matplotlib.collections.QuadMesh at 0x7f07d361c8d0>
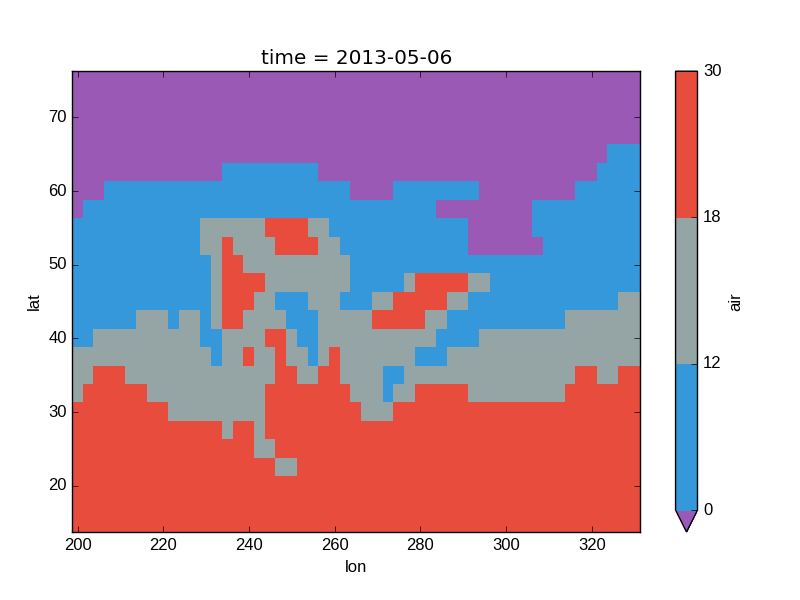
Finally, if you have Seaborn
installed, you can also specify a seaborn color palette to the cmap
argument. Note that levels must be specified with seaborn color palettes
if using imshow or pcolormesh (but not with contour or contourf,
since levels are chosen automatically).
In [46]: air2d.plot(levels=10, cmap='husl')
Out[46]: <matplotlib.collections.QuadMesh at 0x7f07d3510a90>
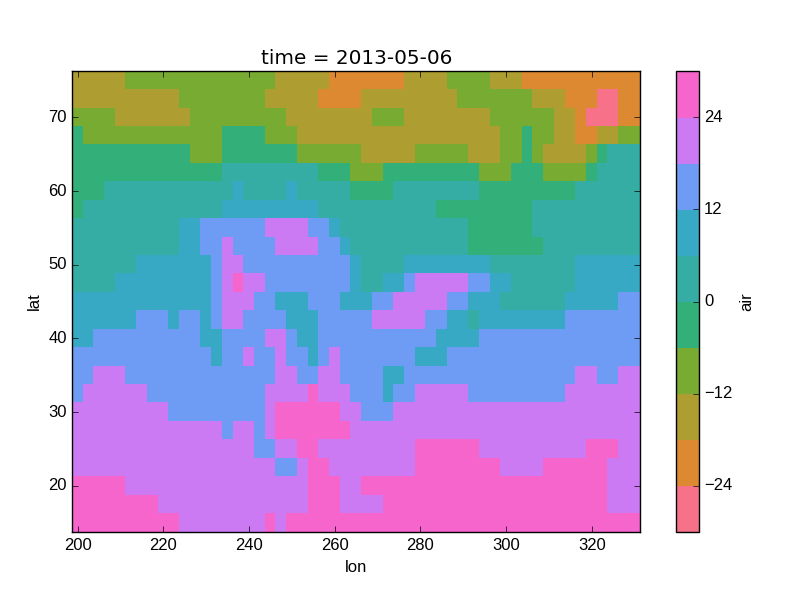
Faceting¶
Faceting here refers to splitting an array along one or two dimensions and plotting each group. xarray’s basic plotting is useful for plotting two dimensional arrays. What about three or four dimensional arrays? That’s where facets become helpful.
Consider the temperature data set. There are 4 observations per day for two years which makes for 2920 values along the time dimension. One way to visualize this data is to make a seperate plot for each time period.
The faceted dimension should not have too many values; faceting on the time dimension will produce 2920 plots. That’s too much to be helpful. To handle this situation try performing an operation that reduces the size of the data in some way. For example, we could compute the average air temperature for each month and reduce the size of this dimension from 2920 -> 12. A simpler way is to just take a slice on that dimension. So let’s use a slice to pick 6 times throughout the first year.
In [47]: t = air.isel(time=slice(0, 365 * 4, 250))
In [48]: t.coords
Out[48]:
Coordinates:
* lat (lat) float32 75.0 72.5 70.0 67.5 65.0 62.5 60.0 57.5 55.0 52.5 ...
* time (time) datetime64[ns] 2013-01-01 2013-03-04T12:00:00 2013-05-06 ...
* lon (lon) float32 200.0 202.5 205.0 207.5 210.0 212.5 215.0 217.5 ...
Simple Example¶
The easiest way to create faceted plots is to pass in row or col
arguments to the xarray plotting methods/functions. This returns a
xarray.plot.FacetGrid object.
In [49]: g_simple = t.plot(x='lon', y='lat', col='time', col_wrap=3)
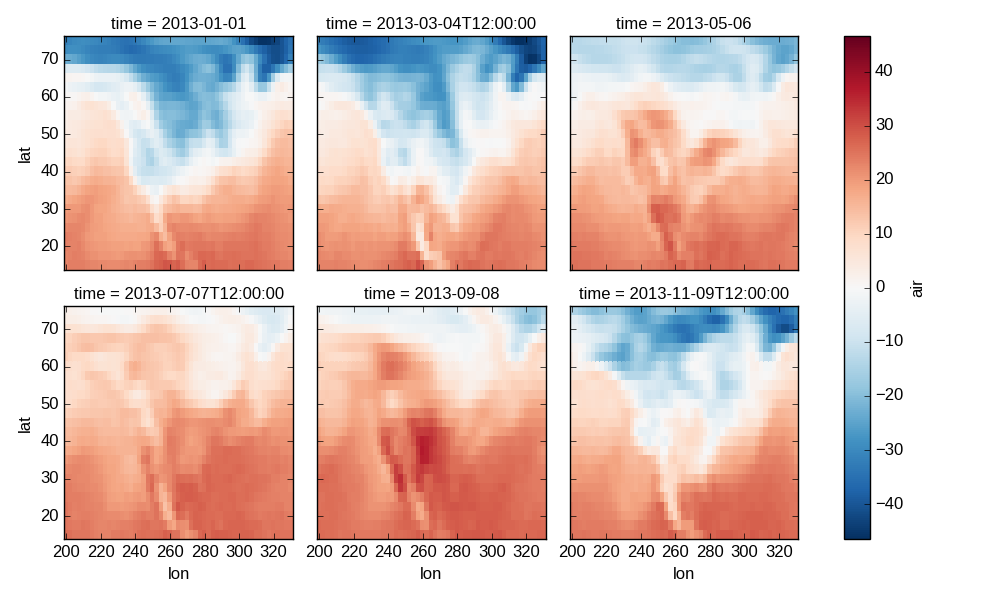
4 dimensional¶
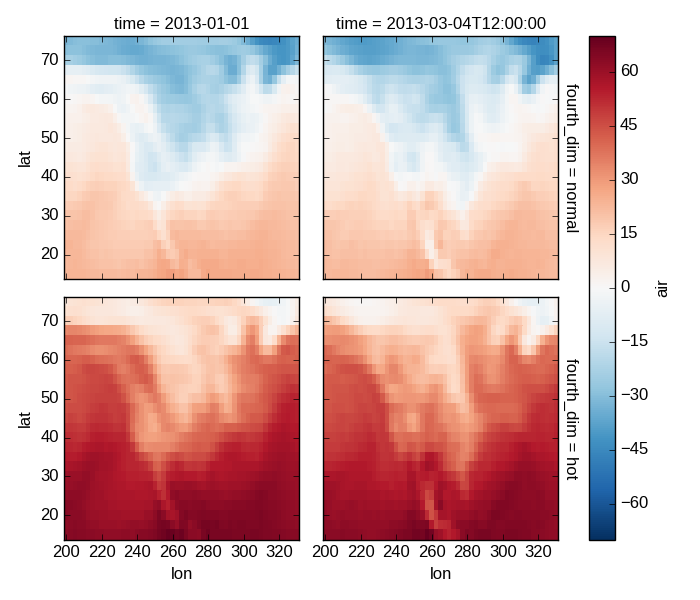
For 4 dimensional arrays we can use the rows and columns of the grids. Here we create a 4 dimensional array by taking the original data and adding a fixed amount. Now we can see how the temperature maps would compare if one were much hotter.
In [50]: t2 = t.isel(time=slice(0, 2))
In [51]: t4d = xr.concat([t2, t2 + 40], pd.Index(['normal', 'hot'], name='fourth_dim'))
# This is a 4d array
In [52]: t4d.coords
Out[52]:
Coordinates:
* lat (lat) float32 75.0 72.5 70.0 67.5 65.0 62.5 60.0 57.5 55.0 ...
* time (time) datetime64[ns] 2013-01-01 2013-03-04T12:00:00
* lon (lon) float32 200.0 202.5 205.0 207.5 210.0 212.5 215.0 ...
* fourth_dim (fourth_dim) object 'normal' 'hot'
In [53]: t4d.plot(x='lon', y='lat', col='time', row='fourth_dim')
Out[53]: <xarray.plot.facetgrid.FacetGrid at 0x7f07d3203710>
Other features¶
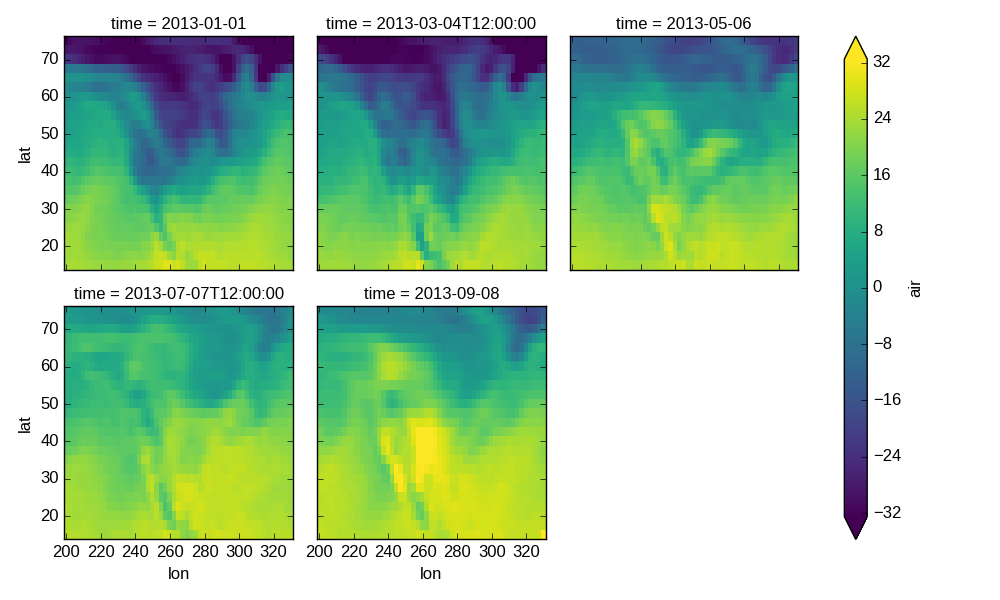
Faceted plotting supports other arguments common to xarray 2d plots.
In [54]: hasoutliers = t.isel(time=slice(0, 5)).copy()
In [55]: hasoutliers[0, 0, 0] = -100
In [56]: hasoutliers[-1, -1, -1] = 400
In [57]: g = hasoutliers.plot.pcolormesh('lon', 'lat', col='time', col_wrap=3,
....: robust=True, cmap='viridis')
....:
FacetGrid Objects¶
xarray.plot.FacetGrid is used to control the behavior of the
multiple plots.
It borrows an API and code from Seaborn.
The structure is contained within the axes and name_dicts
attributes, both 2d Numpy object arrays.
In [58]: g.axes
Out[58]:
array([[<matplotlib.axes._subplots.AxesSubplot object at 0x7f07d2d4bb90>,
<matplotlib.axes._subplots.AxesSubplot object at 0x7f07d2d5fd90>,
<matplotlib.axes._subplots.AxesSubplot object at 0x7f07d2c9d910>],
[<matplotlib.axes._subplots.AxesSubplot object at 0x7f07d2c0c690>,
<matplotlib.axes._subplots.AxesSubplot object at 0x7f07d2b8e490>,
<matplotlib.axes._subplots.AxesSubplot object at 0x7f07d2dfdd10>]], dtype=object)
In [59]: g.name_dicts
Out[59]:
array([[{'time': numpy.datetime64('2013-01-01T00:00:00.000000000')},
{'time': numpy.datetime64('2013-03-04T12:00:00.000000000')},
{'time': numpy.datetime64('2013-05-06T00:00:00.000000000')}],
[{'time': numpy.datetime64('2013-07-07T12:00:00.000000000')},
{'time': numpy.datetime64('2013-09-08T00:00:00.000000000')}, None]], dtype=object)
It’s possible to select the xarray.DataArray or
xarray.Dataset corresponding to the FacetGrid through the
name_dicts.
In [60]: g.data.loc[g.name_dicts[0, 0]]
Out[60]:
<xarray.DataArray 'air' (lat: 25, lon: 53)>
array([[-100. , -30.65, -29.65, ..., -40.35, -37.65, -34.55],
[ -29.35, -28.65, -28.45, ..., -40.35, -37.85, -33.85],
[ -23.15, -23.35, -24.26, ..., -39.95, -36.76, -31.45],
...,
[ 23.45, 23.05, 23.25, ..., 22.25, 21.95, 21.55],
[ 22.75, 23.05, 23.64, ..., 22.75, 22.75, 22.05],
[ 23.14, 23.64, 23.95, ..., 23.75, 23.64, 23.45]])
Coordinates:
* lat (lat) float32 75.0 72.5 70.0 67.5 65.0 62.5 60.0 57.5 55.0 52.5 ...
time datetime64[ns] 2013-01-01
* lon (lon) float32 200.0 202.5 205.0 207.5 210.0 212.5 215.0 217.5 ...
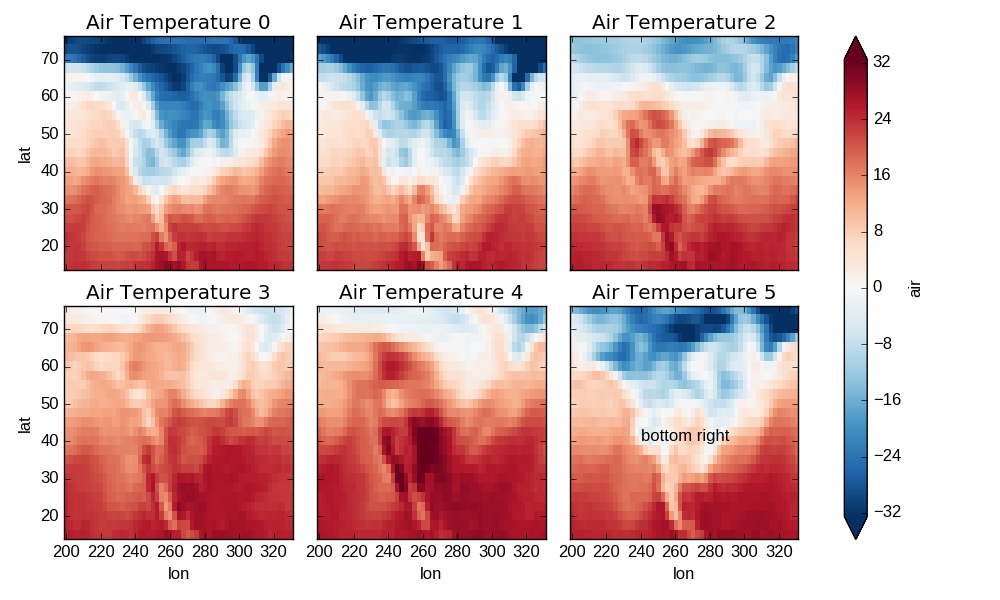
Here is an example of using the lower level API and then modifying the axes after they have been plotted.
In [61]: g = t.plot.imshow('lon', 'lat', col='time', col_wrap=3, robust=True)
In [62]: for i, ax in enumerate(g.axes.flat):
....: ax.set_title('Air Temperature %d' % i)
....:
In [63]: bottomright = g.axes[-1, -1]
In [64]: bottomright.annotate('bottom right', (240, 40))
Out[64]: <matplotlib.text.Annotation at 0x7f07d3323350>
In [65]: plt.show()
TODO: add an example of using the map method to plot dataset variables
(e.g., with plt.quiver).
Maps¶
To follow this section you’ll need to have Cartopy installed and working.
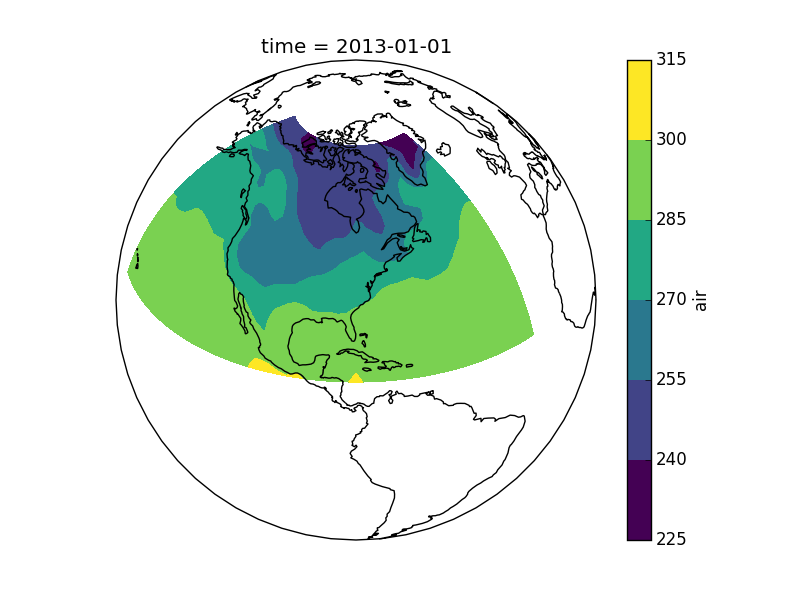
This script will plot the air temperature on a map.
import xarray as xr
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
air = (xr.tutorial
.load_dataset('air_temperature')
.air
.isel(time=0))
ax = plt.axes(projection=ccrs.Orthographic(-80, 35))
ax.set_global()
air.plot.contourf(ax=ax, transform=ccrs.PlateCarree())
ax.coastlines()
plt.savefig('cartopy_example.png')
Here is the resulting image:
Details¶
Ways to Use¶
There are three ways to use the xarray plotting functionality:
- Use
plotas a convenience method for a DataArray. - Access a specific plotting method from the
plotattribute of a DataArray. - Directly from the xarray plot submodule.
These are provided for user convenience; they all call the same code.
In [66]: import xarray.plot as xplt
In [67]: da = xr.DataArray(range(5))
In [68]: fig, axes = plt.subplots(ncols=2, nrows=2)
In [69]: da.plot(ax=axes[0, 0])
Out[69]: [<matplotlib.lines.Line2D at 0x7f07d2734090>]
In [70]: da.plot.line(ax=axes[0, 1])
Out[70]: [<matplotlib.lines.Line2D at 0x7f07d2f52350>]
In [71]: xplt.plot(da, ax=axes[1, 0])
Out[71]: [<matplotlib.lines.Line2D at 0x7f07d2734fd0>]
In [72]: xplt.line(da, ax=axes[1, 1])
Out[72]: [<matplotlib.lines.Line2D at 0x7f07d8d57e90>]
In [73]: plt.tight_layout()
In [74]: plt.show()
Here the output is the same. Since the data is 1 dimensional the line plot was used.
The convenience method xarray.DataArray.plot() dispatches to an appropriate
plotting function based on the dimensions of the DataArray and whether
the coordinates are sorted and uniformly spaced. This table
describes what gets plotted:
| Dimensions | Plotting function |
| 1 | xarray.plot.line() |
| 2 | xarray.plot.pcolormesh() |
| Anything else | xarray.plot.hist() |
Coordinates¶
If you’d like to find out what’s really going on in the coordinate system, read on.
In [75]: a0 = xr.DataArray(np.zeros((4, 3, 2)), dims=('y', 'x', 'z'),
....: name='temperature')
....:
In [76]: a0[0, 0, 0] = 1
In [77]: a = a0.isel(z=0)
In [78]: a
Out[78]:
<xarray.DataArray 'temperature' (y: 4, x: 3)>
array([[ 1., 0., 0.],
[ 0., 0., 0.],
[ 0., 0., 0.],
[ 0., 0., 0.]])
Coordinates:
* y (y) int64 0 1 2 3
* x (x) int64 0 1 2
z int64 0
The plot will produce an image corresponding to the values of the array. Hence the top left pixel will be a different color than the others. Before reading on, you may want to look at the coordinates and think carefully about what the limits, labels, and orientation for each of the axes should be.
In [79]: a.plot()
Out[79]: <matplotlib.collections.QuadMesh at 0x7f07d3468910>
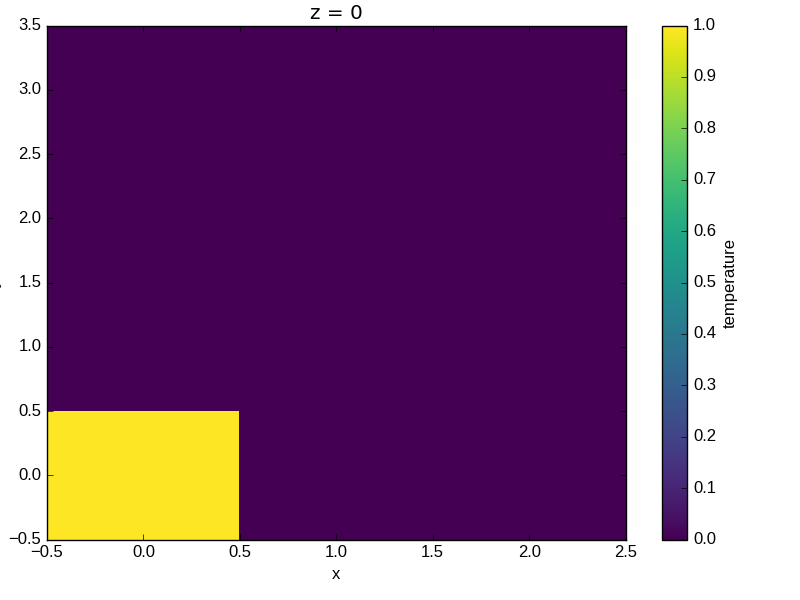
It may seem strange that the values on the y axis are decreasing with -0.5 on the top. This is because the pixels are centered over their coordinates, and the axis labels and ranges correspond to the values of the coordinates.